Since the original release of the specification for the Next Generation File Format (NGFF) OME-Zarr, Glencoe Software has been dedicated to building implementations of these concepts. Now has come the time to optimize for the particular requirements of bio-imaging data and the ways we work with it.
Glencoe Software’s work with OME-Zarr tooling includes applications and libraries for writing (bioformats2raw), reading (raw2ometiff), analyzing (OMERO Segmentation Connector) and managing (OMERO Plus) data in this format. Throughout these efforts, we have relied on third party libraries as a member of a larger ecosystem interacting with the array (Zarr) and storage (S3) technologies relevant for cloud-native image data management. While this strategy is both practical and collaborative, it also risks missing the opportunity to specialize these implementations for the realities of bio-imaging data. Our next release of OMERO Plus includes new optimizations described below, and we have ideas for further improvements to be realized over the remainder of this year.
Optimizations included in the upcoming OMERO Plus release:
Optimization of s3fs
Based on our evaluation of the original s3fs implementation relied upon by OMERO Plus, there remained a number of patterns of communication with S3 that reflected operations which are reasonable when working with a local file system, but incredibly expensive and unnecessary when working with object storage in read-only mode. Further, there were understandably no optimizations that took advantage of the predictable layout of the OME-Zarr format. In particular, OME-Zarr has predictable chunk sizes and the expectation that all of the objects within the parent .zarr prefix have the same access permissions. The result was repetitive requests for the same objects and unnecessary object size and authorization checks. In our reimplementation of this functionality, we have removed these repetitive and unnecessary requests, with the result that there is a single request to S3 made per channel and per chunk for any tile to be rendered in the viewer.
Parallel requests for RGB data
RGB (Brightfield) data, often of whole slide images, is a visualization-heavy workflow where latency of the viewer matters substantially. Although gamma and color correction may be performed prior to review, the visualization process most often considers all three channels as a unit, unlike fluorescence where channel visibility, min/max, and pseudocolors are changed often. We can therefore optimize the reading of this data as 3 parallel requests, covering the Red, Green and Blue channels.
Caching of rendering metadata
Image-level rendering metadata can be cached, such that subsequent tiles in the viewer are even faster to appear than the first.
Results
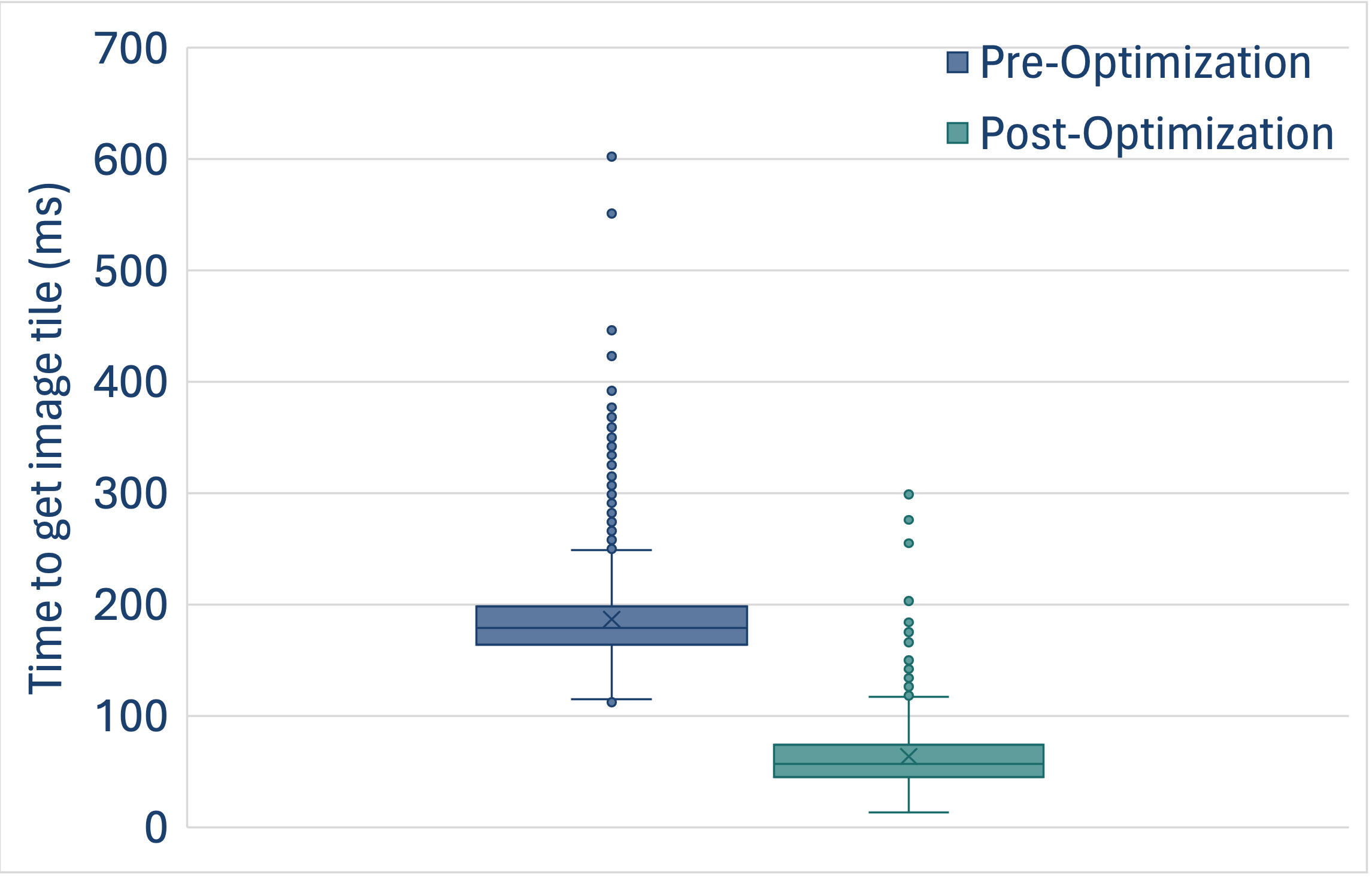
The plot demonstrates the improvements in the time to read a single tile (in a manner matching how OMERO Plus reads this data) with and without the above changes. This analysis was performed using Glencoe’s benchmarking tool, which is available for evaluation of various image data storage backends as our customers choose the appropriate storage architecture for their OMERO Plus use cases.
Areas of future work:
Generalized parallel requests
OMERO and OMERO Plus currently render image data in a serial, per-channel fashion. As described above, working with S3 enables higher parallelism of data retrieval, such that multiple channels could be read at once. Changes will be proposed to the core of OMERO and OMERO Plus to enable such parallel rendering activity.
Further optimizations for RGB data
Optimizations for RGB (Brightfield) data can be further considered - including single, interleaved 3-channel chunks, with added support for lossy compression. These changes would impact OME-Zarr specification overall, so they must be done with community engagement.
Conclusion
While we have diverged from existing third party libraries to forge a path necessary for our customers’ success, this is not an indication that we alone should learn and benefit from these changes. We have made the decision to make the components for OME-Zarr reading (via S3) in OMERO Plus public, as we hope these serve as an example of the requirements for working with bio-imaging data. We are glad to widen the discussion on how we prioritize work on these basic data and storage libraries to benefit the wider bio-imaging community.
Questions or comments? We would love to hear from you. Reach out to us at info@glencoesoftware.com.
